Welcome to DeepCalpain
DeepCalpain is a web server developed for understanding the enzyme-specific cleavage for calpains including m-calpain and μ-calpain. Based on deep learning method, the predictor achieved promising performances. Four bright spots of DeepCalpain software are summarized below: i) four kinds of protein sequence features are modeled independently and then merged into one model; ii) particle swarm optimizer algorithm is used to optimize the hyperparameters; iii) the protein-protein interaction network and colocalization information are provided and visulized in the result page; iv) three fundamental properties of protein sequence are visualized in the result page, including disorder, surface accessibility and secondary structure.
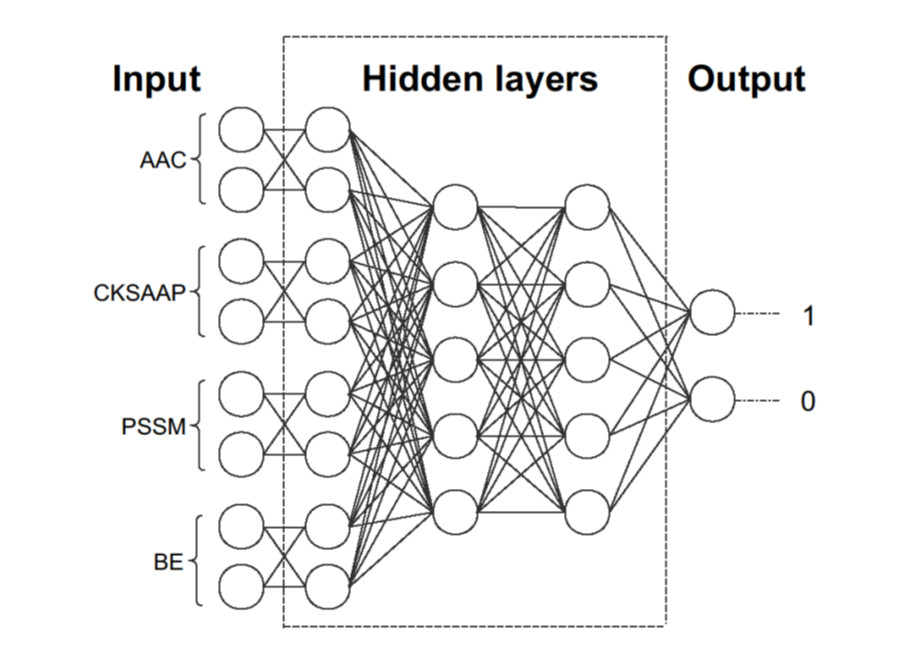
Deep neural network for prediction of calpain-specific cleavage
• Four sequence features of the calpain cleavage sites are extracted to build models.
• Four models are combined into one full connection layer.
• Deep-learning method is used to predict calpain-specific cleavage sites.
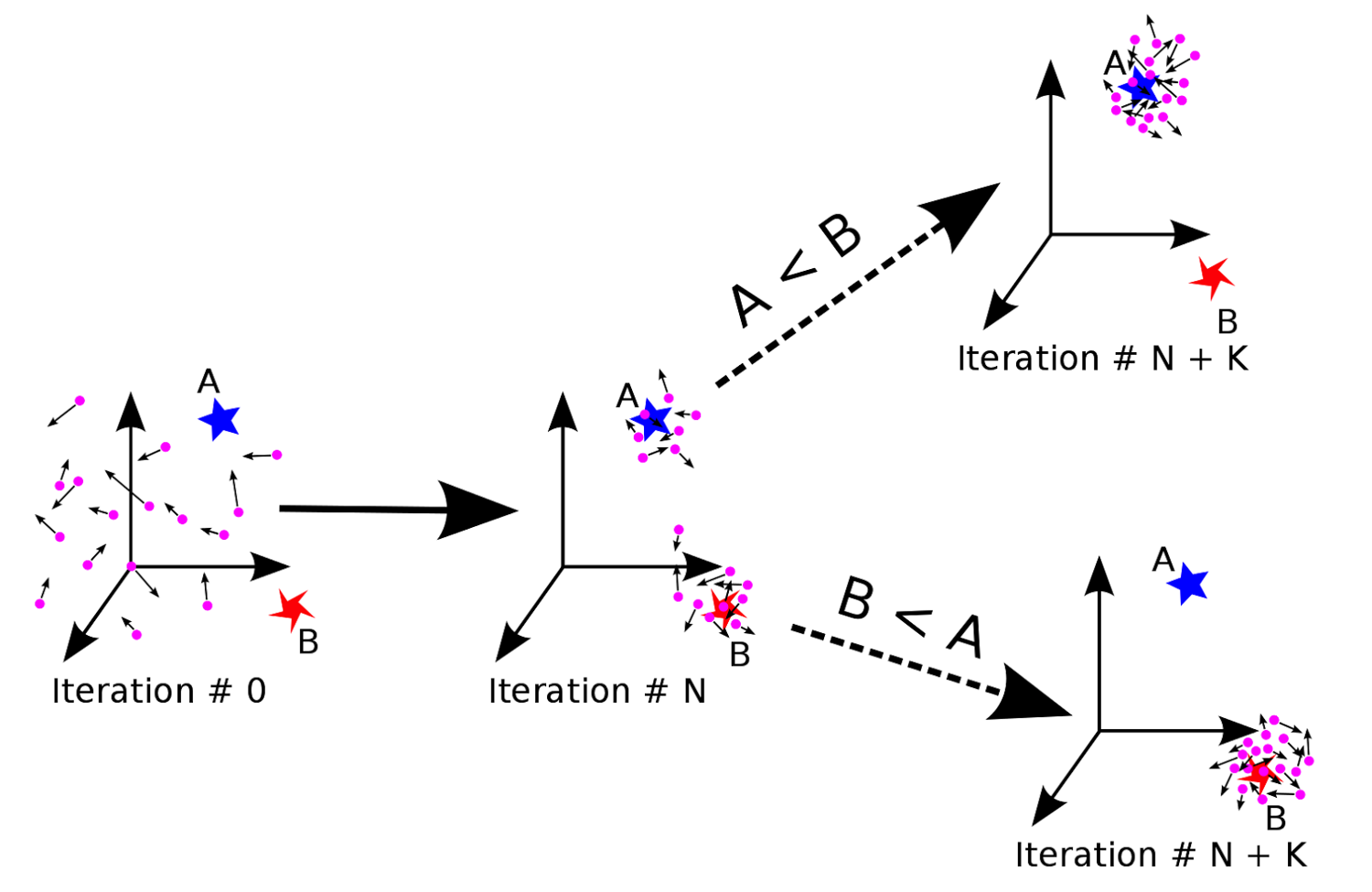
Optimize the hyperparameters of DeepCalpain
• Deep neural network contains hyperparameters such as learning rates, activation functions, dropout rates, etc.
• Taking both efficiency and effectiveness into account, particle swarm optimizer (PSO) is applyed to optimize the hyperparameters.
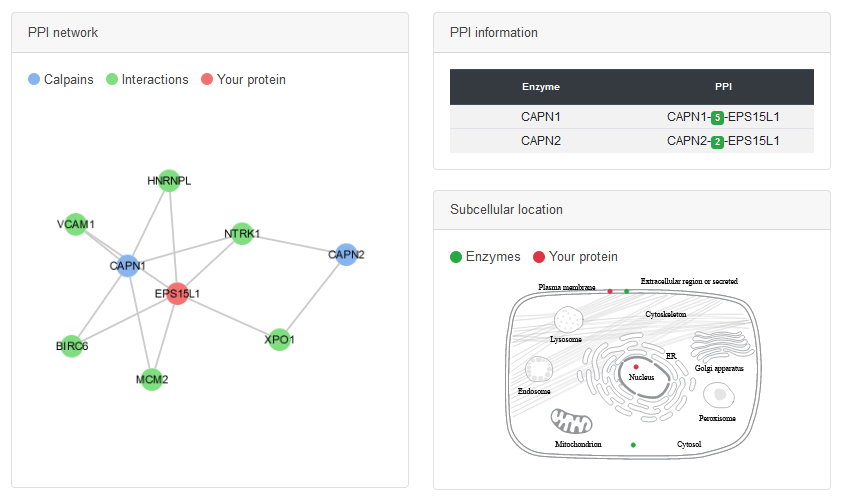
PPI network
• The protein-protein interaction (PPI) data are collected from several databases.
• The cell map shows the colocalization between calpain and the substrate.
• In addition to the direct PPI, the proteins connected to calpain via scaffold proteins are also shown.
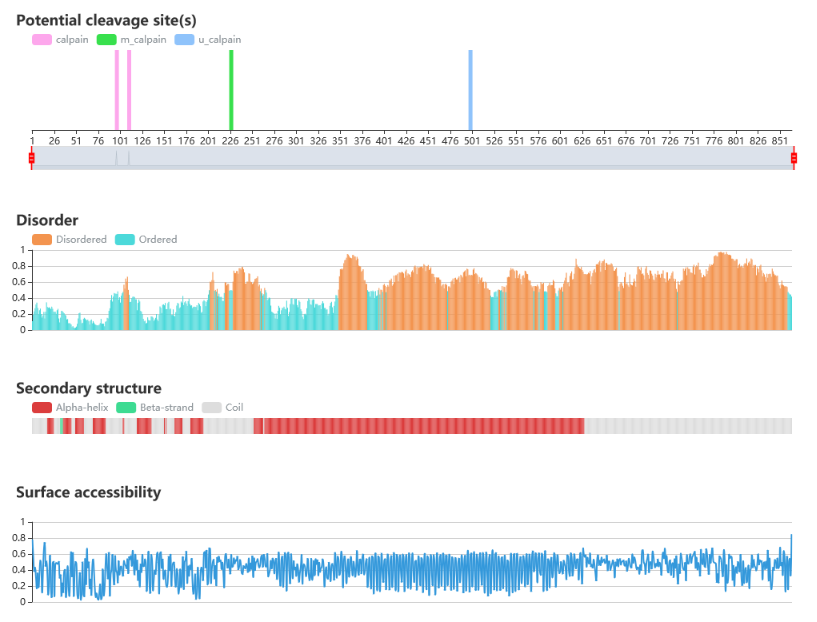
Protein sequence and structure properties
• IUPred is used to predict the disorder region.
• NetSurfP is used to predict the surface accessibility and secondary structure.
• The potential cleavage sites of the query sequence are indicated.
